Tami McDonald was the recipient of the Mason Hale Award from the International Association for Lichenology (IAL) for the best Ph.D. thesis in lichenology completed between August 1, 2011 and January, 31, 2014. The award was announced at the 10th International Mycological Congress (IMC10) in Bangkok, Thailand. Thesis title: Genomic insights into the evolution and development of the lichen symbiosis: Cladonia grayi as a model lichen.
Category: News
2014-08: Congratulations Ko-Hsuan (Koko) Chen – Best poster award at IMC10.
Ko-Hsuan (Koko) Chen (Ph.D. student) was one of the recipients of the best poster awards at the 10th International Mycological Congress (IMC10), Bangkok, Thailand. The title of her poster was “Phylogenetic analyses of eurotiomyceteous endophytes reveal their close affinities to Chaetothyriales, Eurotiales and a new order – Phaeomoniellales.”
2014-06: Congratulations Tami McDonald – position at St. Catherine University.
Tami McDonald, who is a Ph.D. alumni of the Lutzoni lab, and currently a postdoctoral research associate at the University of Minnesota, Twin Cities, accepted a tenure-track, Assistant Professor, position in the Department of Biology at St. Catherine University, St. Paul, Minnesota. Her starting date at St. Kate’s is August 18, 2014.
2013-05: Congratulations Ester Gaya – position at Kew Gardens.
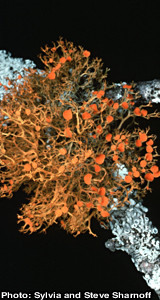 May 2015: Congratulations Ester Gaya
May 2015: Congratulations Ester Gaya
Ester Gaya, who has been pursuing postdoctoral research in our lab as part of an NSF funded phylogenetic study of the order Teloschistales, accepted a permanent position as Senior Researcher in Mycology, at Kew Royal Botanic Gardens, London, UK. Her starting date at Kew is July 1, 2013
2013-05: Congratulations Ryoko Oono – position at UCSB.
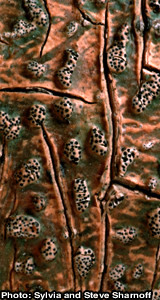 |
May 2013: Congratulations Ryoko OonoRyoko Oono, who is currently pursuing postdoctoral research in our lab as part of the Molecular Mycology and Pathogenesis Training Program (MMPTP), accepted a tenure-track, Assistant Professor, position in the department of Ecology, Evolution and Marine Biology at UCSB. Her starting date at UCSB is Jan. 1, 2014. |
2013-03: Congratulations Kathryn Picard.
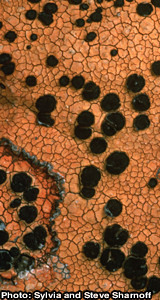 March 2013: Congratulations Kathryn Picard
March 2013: Congratulations Kathryn Picard
Congratulations, Kathryn Picard, for being awarded a NSF Doctoral Dissertation Improvement Grant!
2006-12: Farewell, Frank Kauff, Postdoctoral Research Associate.
 |
December 2006 – Farewell, Frank Kauff, Postdoctoral Research Associate.The members of the Lutzoni lab send their best wishes to Frank Kauff, a postdoctoral research associate who has worked with us for four years. He begins a new position on December 15, 2006, as a Junior Professor in Molecular Phylogenetics of lower plants at the University of Kaiserslautern, Germany. Congratulations and good luck to Frank as he moves on to the next stage. |
2007-01: Farewell, Valerie Reeb, Graduate Student and Postdoctoral Research Associate.
 |
January 2007 – Farewell, Valerie Reeb, Graduate Student and Postdoctoral Research Associate.It is time to bid farewell to our own Valerie Reeb who has been a member of the Lutzoni lab for many years as a technician, graduate student and a postdoctoral research associate. We would like to congratulate her on obtaining a two year post-doctoral position at the University of Iowa with Debashish Bhattacharya. She will be working on the evolution and phylogeny of microbial eukaryotes. The project, part of an AToL (Assembling the Tree of Life) grant, consists of sequencing 9 genes for 200 taxa in order to elucidate the eukaryotic tree of life and address specific hypotheses regarding eukaryotic evolution. Congratulations and good luck to our first PhD! |
2007-05: Farewell, Samantha Hill, Undergraduate.
 |
May 2007 – Farewell, Samantha Hill, Undergraduate.Samantha has been with us for many years in the lab keeping everything running smoothly from week to week. From pouring culture plates to autoclaving glassware, she has always provided us with our research needs. We will miss her as she moves on to bigger and better things! |
2007-10: Congratulations to Brendan Hodkinson for passing his prelims.
 |
October 2007 –Congratulations to Brendan Hodkinson for passing his prelims! |